Super Charged Python
In my first blog post, I explained how to perform X-Ray imaging. One problem with a pure Python forward model is it’s slow. Therefore, I am going to introduce Numba in this blog post a way to accelerate Python code.
Profiling
One way to determine why code is slow is by profiling it. One of my recommended resources to profile Python code is to use line_profiler. A line profiler will measure the time to execute each line of code and the # of calls to that line. line_profiler can be used by placing a @profile decorator above the functions to profile and running the Python code with the command kernprof -l *NAME*.py. Bellow is an example of running line_profiler on the X-ray algorithm. Another line profiler to check out is VTune offered by Intel.
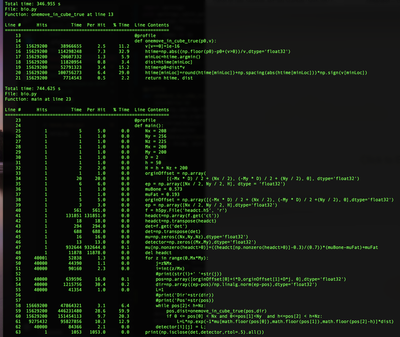
One result from the line profile report is many function calls have a high # hits(# of calls) with little execution time per call(lines 49-62). Another nice feature of line profiler is the percent time which highlights the one move in cube function is taking a lot of time. A way to improve the speed of this code is by either A) Decreasing the amount of time that each function call takes or B) Creating portions of the code that can run in parallel. This blog post will focus on A using a library called Numba to make each function faster.
Numba
Example Numba Function
@jit(nopython=True, cache=True)
def square_log(x):
return x*x+np.log(x)
square_log(np.arange(5000))
Numba is a just in time compiler(complies code when called) that can speed up numerical code that uses some constrained set of NumPy functions and Python features. The ray tracing code is a great use case for this as all the data structures and data manipulation is in NumPy. One of the easiest ways to use Numba is through the decorator @jit. Putting this decorator on top of a function causes Numba to compile that function using LLVM and replace it with a possibly faster version. @jit can take various arguments including cache to cache the compiled function, parallel which will try to parallelize any part of the code and nopython which forces nopython mode. Numba has two different modes, object mode and nopython mode. nopython mode forces all the data types in Numba to be represented as numeric types rather than Python objects. Python object mode is slower than nopython mode as Numba is constrained to optimize numerical code rather than pure Python code. One example of using Numba in the X-Ray algorithm is placing @jit above the main loop and one move in cube function.
What is Numba doing?
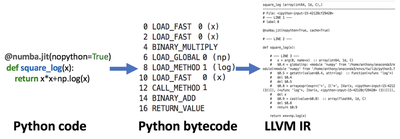

Numba makes code faster by compiling it and running an optimizer on it to make it faster. Compiling Python Code!? Yup, you heard me right, Numba uses LLVM to take Python’s bytecode and turn it into LLVM IR. LLVM IR is an intermediate representation used for any language which LLVM then performs various optimization passes(more info). After running the @jit decorator one way to view the LLVM IR produced is through the inspect_types method. Additionally to force Numba to use nopython mode the developed code must use the supported Python Features and NumPy Functions, otherwise, the slower object mode will be used.
How much faster is Numba??
Bellow is an example of the difference in speed between Numba and pure Python. Numba is able to accelerate the code by ~100x compared to the pure Python code. The main reason behind this is Numba is restricting the dynamicity of Python to static types and optimizing various calls to take advantage of this fact. For further information on the dynamicity slowdown in Python see this article from the 2018 Python Language Summit.
For more information on Numba: see the documentation.
That wraps up this blog post, next time I will review one more way to speed up the X-Ray solver using a GPU with Numba.